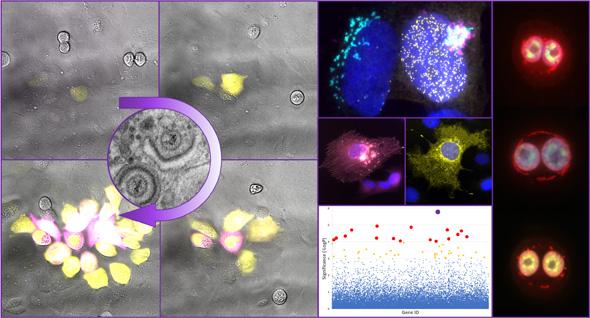
Our Research
Our laboratory studies cellular mechanisms utilised and modified by viruses during their replication. We currently study viruses from important families of human and veterinary pathogens: herpesviruses, polyomaviruses, and bunyaviruses.
Herpesviruses are large and complex enveloped dsDNA viruses that are ubiquitous pathogens of vertebrates and establish life-long latent infections in their hosts. Infections by the nine known human herpesviruses are associated with many serious diseases including certain lymphomas and life-threatening conditions in immuno-compromised patients. Herpesviruses encode a large number of multi-functional proteins that cause dramatic or subtle modifications of host cells enabling the evasion of antiviral responses and efficient production of new virions.
Polyomaviruses are small non-enveloped dsDNA viruses that are ubiquitous pathogens in humans and can establish life-long persistent infections with regular asymptomatic virus shedding. While polyomavirus infections are generally benign in immuno-competent hosts they are known to cause serious disease in immuno-compromised patients including haemorrhagic cystitis, polyomavirus-associated nephropathy, Merkel cell carcinoma, and progressive multifocal leukoencephalopathy.
Bunyaviruses are enveloped segmented negative-strand RNA viruses that are commonly transmitted by arthropods and includes many neglected tropical disease pathogens such as Rift Valley fever virus, Crimean-Congo haemorrhagic fever virus, and Oropouche virus. In collaboration with colleagues at São Paulo University we are investigating virus-cell interactions involved in the morphogenesis and egress of Oropouche virus, one of the most common arboviruses that infects humans in South America.
Our current projects
- Host cell modulation by herpesvirus proteins
- Mechanisms of polyomavirus egress
- Viral and cellular mechanisms of herpesvirus cell-to-cell spread
- Identification of “druggable” host factors required for polyomavirus replication
- Advanced imaging techniques to analyse virus assembly and spread
- Bunyavirus-host cell interactions
Recent Publications
Benedyk TH, Muenzner J, Connor V, Han Y, Brown K, Wijesinghe KJ, Zhuang Y, Colaco S, Stoll GA, Tutt OS, Svobodova S, Svergun DI, Bryant NA, Deane JE, Firth AE, Jeffries CM, Crump CM†, Graham SC† (2021) pUL21 is a viral phosphatase adaptor that promotes herpes simplex virus replication and spread. PLoS Pathog. 2021 Aug 16;17(8):e1009824. doi: 10.1371/journal.ppat.1009824
Scherer KM, Manton JD, Soh TK, Mascheroni L, Connor V, Crump CM†, Kaminski CF† (2020) A fluorescent reporter system enables spatiotemporal analysis of host cell modification during herpes simplex virus-1 replication. J Biol Chem. 2020 Dec 30;296:100236
Soh TK*, Davies CTR*, Muenzner J*, Hunter, LM, Barrow, HG, Connor V, Bouton C, Smith C, Emmott E, Antrobus R, Graham SC†, Weekes MP†, Crump CM† (2020) Whole-cell and plasma membrane proteomics of herpes simplex virus-1 infection reveals cell surface remodelling via pUL56-mediated GOPC degradation. Cell Rep 33: 108235
Caller LG, Davies CTR, Antrobus R, Lehner PJ, Weekes MP†, Crump CM† (2019) Temporal Proteomic Analysis of BK Polyomavirus Infection Reveals Virus-Induced G2 Arrest and Highly Effective Evasion of Innate Immune Sensing J Virol 93:e00595-19. doi: 10.1128/JVI.00595-19
Albecka A, Owen DJ, Ivanova L, Brun J, Liman R, Davies L, Ahmed MF, Colaco S, Hollinshead M, Graham SC†, Crump CM† (2017) Dual Function of the pUL7-pUL51 Tegument Protein Complex in Herpes Simplex Virus 1 Infection. J Virol. 2017 Jan 3;91(2). pii: e02196-16. doi: 10.1128/JVI.02196-16
Dr Colin Crump
Principal Investigator
Group Members


