Our Research
As we examine the world around us, we see a story of evolutionary success: every organism now living has developed characteristics that enable it to survive better than similar organisms that died out. By examining genetic sequences, it is possible to look for genetic signatures of such survival characteristics. We develop and apply mathematical and computational techniques to examine the sequences of RNA viruses, looking for conserved regions of the genome that suggest missing viruses that have died out. In this way, we aim to find features of these viruses critical to their lifecycles, which can be characterised experimentally. The ultimate aim, in keeping with our clinical orientation, is to find targets for new antiviral drugs. Our work is at the interface between mathematics and molecular virology.
In addition to our main focus on viral genetics, we undertake work aiming to improve the use of diagnostics in clinical laboratories, and modelling of the dynamics of viral disease to provide answers to policy questions on testing and treatment.
Current Topics
- Computational genetics of RNA viruses
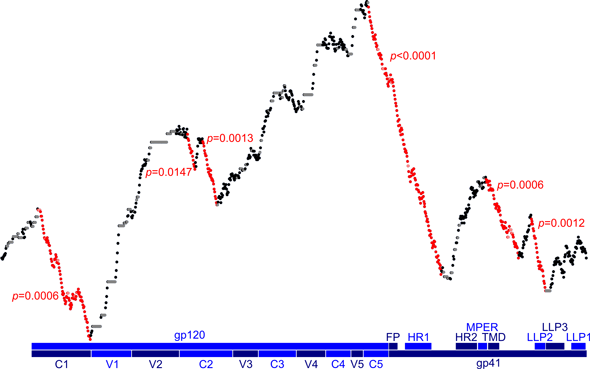
- HIV-1 structure and function
- SARS-CoV-2 structure and function
- Flavivirus structure and function
- Properties of clinical diagnostic tests
- Testing and treatment policy for viral disease
Key publications
Julia R. Gog, Andrew M. L. Lever, Jordan P. Skittrall. A new method for detecting signal regions in ordered sequences of real numbers, and application to viral genomic data. PLoS ONE 13(4): e0195763. doi:10.1371/journal.pone.0195763
PMID: 29652903
Jordan P. Skittrall, Carin K. Ingemarsdotter, Julia R. Gog, Andrew M. L. Lever. A scale-free analysis of the HIV-1 genome demonstrates multiple conserved regions of structural and functional importance. PLoS Computational Biology 15(9):e1007345. doi:10.1371/journal.pcbi.1007345 PMID: 31545786
Jordan P. Skittrall, Michael Wilson, Anna A. Smielewska, Surendra Parmar, Mary D. Fortune, Dominic Sparkes, Martin D. Curran, Hongyi Zhang, Hamid Jalal. Specificity and positive predictive value of SARS-CoV-2 nucleic acid amplification testing in a low-prevalence setting. Clinical Microbiolgy and Infection (2020). doi:10.1016/j.cmi.2020.10.003 PMID: 33068757
Jordan P. Skittrall SARS-CoV-2 screening: effectiveness and risk of increasing transmission. Journal of the Royal Society Interface, 2021 18(180) 20210164. doi:10.1098/rsif.2021.0164 PMID: 34283945


