Research
 Our group is interested in defining the mechanisms by which chromatin modifications function to regulate cellular processes. Our attention is focused on a set of enzymes (acetylases, deacetylases, methylases and kinases), which regulate transcription by covalently modifying histones. We would like to understand what biological processes these enzymes control and the precise role of each modification on chromatin dynamics. In addition, a number of chromatin modifying enzymes have been implicated in the genesis of cancer so we are dissecting how these pathways missregulated in cancer cells.
Our group is interested in defining the mechanisms by which chromatin modifications function to regulate cellular processes. Our attention is focused on a set of enzymes (acetylases, deacetylases, methylases and kinases), which regulate transcription by covalently modifying histones. We would like to understand what biological processes these enzymes control and the precise role of each modification on chromatin dynamics. In addition, a number of chromatin modifying enzymes have been implicated in the genesis of cancer so we are dissecting how these pathways missregulated in cancer cells.
Histones are very highly modified. Despite their abundance, we believe that more modifications are likely to exist on histones. This complexity is probably necessary because histones integrate many signalling pathways with biological processes involving DNA manipulation. We are taking a number of complementary approaches to characterise the function of chromatin modifications. We use yeast as a model system whenever possible to define pathways. We use human cells to characterise function in higher organisms and probe connections to cancer. Mechanistic analysis of modifications is carried out using recombinantly assembled nucleosomes that are modified at specific residues.
- Group Members:
Andrej Alendar, Paulo Amaral, Andrew Bannister, Isaia Barbieri, Ester Cannizzaro, Harvey Che, Alistair Cook, Sri Lestari, Nikki Mann, Valentina Migliori, Gonzalo Millan Zambranom, Sa Robson, Helena Santos-Rosa, Meike Wiese
Publications
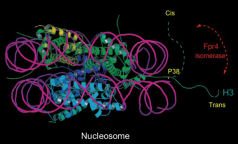
- Tessarz P, Kouzarides T. (2014) Histone core modifications regulating nucleosome structure and dynamics. Nat Rev Mol Cell Biol 15(11):703-708
- Maria Christophorou , Gonçalo Castelo-Branco , Richard Halley-Stott , Clara Slade Oliveira , Remco Loos , Paul Bertone , Jose Silva , Magdalena Zernicka-Goetz , Michael Nielsen , John Gurdon , Aliaksandra Radzisheuskaya , Kerri Mowen & Tony Kouzarides. (2014). Citrullination regulates pluripotency and H1 linker histone binding to chromatin. Nature. 507(7490), 104-108.
- Peter Tessarz, Helena Santos-Rosa, Sam C. Robson, Kathrine B. Sylvestersen,Christopher J Nelson, Michael L. Nielsen & Tony Kouzarides. (2014) Glutamine methylation in Histone H2A is an RNA Polymerase I dedicated modification. Nature. 505(7484), 564-568.
- Blerta Xhemalce, Sam Robson & Tony Kouzarides. (2012) Human RNA Methyltransferase BCDIN3D Regulates MicroRNA Processing. Cell. 151(2), 278-288.
- Mark Dawson, Rab Prinjha, Antje Dittmann, George Giotopoulos, Marcus Bantscheff, Wai-In Chan, Samuel C. Robson, Chun-wa Chung, Carsten Hopf, Mikhail M. Savitski, Carola Huthmacher, Emma Gudgin, Dave Lugo, Soren Beinke, Trevor D. Chapman3, Emma Roberts, Peter Soden3, Kurt Auger, Olivier Mirguet, Konstanze Doehner, Ruud Delwel, Alan Burnett, Phillip Jeffrey, Gerard Drewes, Kevin Lee, Brian Huntly & Tony Kouzarides. (2011) Inhibition of BET recruitment to chromatin as an effective treatment for MLL-fusion leukaemia. Nature. 478(7370), 529-533.
- Till Bartke, Michiel Vermeulen, Blerta Xhemalce, Samuel C. Robson, Matthias Mann, and Tony Kouzarides. (2010). Nucleosome-interacting Proteins Regulated by DNA and Histone Methylation. Cell, 143: 470 – 84


