Research
The lab’s research projects are focused around the innate immune responses to virus infection in three main areas:
1) The innate immune response to viral nucleic acids.
Nucleic acids are powerful immunostimulatory agents. Sensing of damaged or mis-localised nucleic acids by pattern recognition receptors underlies multiple pathologies and is especially important in the context of anti-viral immunity.
Sensing of viral DNA in mammalian cells:
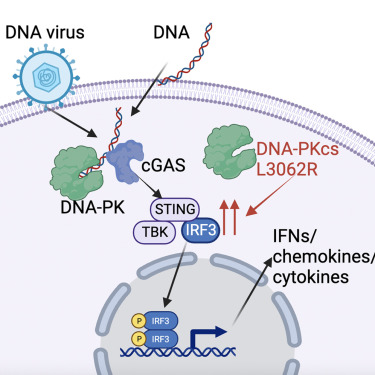
During DNA virus infection, the ability of our cells to sense viral genomic DNA it critical for mounting type one and type three interferon responses. Leveraging our expertise in poxvirology, a family of large DNA viruses which replicate their double stranded DNA genome in the cytoplasm, we discovered that mouse and human cells require DNA-dependent protein kinase (DNA-PK) for this process. DNA-PK binds to cytoplasmic DNA and activates the cGAS/STING pathway that drives the IRF-3 dependent production of interferons and cytokines in response to viral DNA. This pathway is required for mammalian cells to sense and respond to DNA viruses like herpesviruses and poxviruses. We also discovered that patients who have specific mutations in DNA-PKcs (the catalytic subunit of DNA-PK), can have enhanced STING signalling and IFN-I production, showing that this pathway functions in people and can lead to autoinflammatory pathology.
Sensing of viral RNA in mammalian cells:
The receptors that respond to viral RNA, such as RIG-I, MDA5 and TLR3, are equally important for triggering mammalian interferon responses to RNA virus infections. We are using our understanding of cell signalling at the molecular, biochemical and cell biological level to determine the mechanisms by with RNA sensing receptors work. For example we have defined the contribution of the linear ubiquitin chain assembly complex (LUBAC) and M1-linked ubiquitin chains to anti-viral immunity via the regulation of RNA sensing pattern recognition receptors, TLR3 and RIG-I in the context of influenza, Zika and Sendai virus infection. For example, cpdm mice, that lack SHARPIN are more susceptible to influenza infection:
We also found differing roles for SHARPIN and HOIP (the catalytic component of LUBAC) in regulation of RIG-I signalling, with HOIP and HOIL-1 being required for antiviral RIG-I signalling but SHARPIN not, despite it being recruited to the RIG-I signalling complex via NEMO:
2) Evolution of poxvirus host/pathogen interactions
In to understand how nucleic acid sensing mechanisms evolved to drive anti-viral immunity we are taking a comparative immunology approach. We are defining nucleic acid sensing mechanisms in multiple species including humans, mice and birds and, in parallel, defining how poxviruses subvert these pathways to optimise their replicative niches and driving evolution of host defence mechanisms. We have show, for example, that the cGAS/STING pathway functions in chickens and that this is subverted by Fowlpox virus, a poxvirus that infects chickens, and that the mammalian CpG DNA sensing TLR9 pathway is functionally replaced in birds by TLR21:
3) Vaccinology
There are many fundamental questions about the development of immunological memory which remain poorly understood. The initial detection of a vaccine vector by the innate immune system has long lasting consequences for the development of memory responses. We use vaccinia virus, the vaccine that was developed to eradicate smallpox to help understand these fundamental mechanisms of vaccinology. For example we have developed a panel of mutant viruses which have differential impacts on the development of lymphocyte memory. We have defined the impact of removing individual and multiple immunomodulatory proteins from vaccinia virus on vaccination responses and the generation of CD8 T cell memory
We are also working to understand the mechanisms by with vaccinia virus can generate such powerful memory immunity. We have recently discovered that vaccinia virus sabotages dendritic cell function and enters the draining lymph node to drive CD4 T cell activation, and that the pathology at the site of vaccination is almost completely dependent on opportunistic bacterial co-infection.
Publications
| DNA-PKcs is required for cGAS/STING-dependent viral DNA sensing in human cells Dayana B Hristova, Marisa Oliveira, Emma Wagner, Alan Melcher, Kevin J Harrington, Alexandre Belot, Brian J Ferguson iScience, in press |
2023 |
| Contrasting functions of ATP hydrolysis by MDA5 and LGP2 in viral RNA sensing Rahul Singh, Yuan Wu, Alba Herrero del Valle, Kendra E Leigh, Mark TK Cheng, Brian J Ferguson, Yorgo Modis bioRxiv, 2023.05. 25.542247 |
2023 |
| Mechanisms of type I interferon production by chicken TLR21 Rodrigo Guabiraba, Damaris Ribeiro Rodrigues, Paul T Manna, Mélanie Chollot, Vincent Saint-Martin, Sascha Trapp, Marisa Oliveira, Clare E Bryant, Brian J Ferguson Developmental & Comparative Immunology 151, 105093 |
2023 |
| LUBAC is required for RIG-I sensing of RNA viruses Helena C Teague, Charlotte Lefevre, Eva Rieser, Diego de Miguel, Daniel Patricio, Marisa Oliveira, Daniel S Mansur, Nerea Irigoyen, Henning Walczak, Brian J Ferguson Cell Death & Differentiation, 1-12 |
2023 |
| Transcriptional reprogramming of natural killer cells by vaccinia virus shows both distinct and conserved features with mCMV Delphine M Depierreux, Geoffrey L Smith, Brian J Ferguson Frontiers in Immunology 14, e1093381 |
2023 |
| The gut microbiota and its metabolite butyrate shape metabolism and antiviral immunity along the gut-lung axis in the chicken Rodrigo Guabiraba, Vincent Saint-Martin, Vanaique Guillory, Melanie Chollot, Isabelle Fleurot, Emmanuel Kut, Ferdinand Roesch, Ignacio Caballero, Emmanuelle Helloin, Emilie Chambellon, Brian Ferguson, Philippe Velge, Florent Kempf, Sascha Trapp ResearchSquare Pre-print |
2023 |
| PKR-mediated stress response enhances dengue and Zika virus replication Taissa Ricciardi-Jorge, Edroaldo Lummertz da Rocha, Edgar Gonzalez-Kozlova, Gabriela Flavia Rodrigues-Luiz, Brian J Ferguson, Trevor Sweeney, Nerea Irigoyen, Daniel Santos Mansur Mbio 14 (5), e00934-23 |
2023 |
| Suppression of innate immunity by the vaccinia virus protein N1 promotes skin microbiota expansion and increased immune infiltration following vaccination Evgeniya Shmeleva, Daneil Syafiq, Anna Moldoveanu, Brian J Ferguson, Geoffrey L Smith Journal of General Virology 103 (11) |
2022 |
| DNA-PKcs restricts Zika virus spreading and is required for effective antiviral response Daniel de Oliveira Patricio, Greicy Brisa Malaquias Dias, Lucilene Wildner Granella, Ben Trigg, Helena Claire Teague, Dina Bittencourt, André Báfica, Alfeu Zanotto-Filho, Brian Ferguson, Daniel Santos Mansur Frontiers in Immunology 13, 1042463 |
2022 |
| Smallpox vaccination induces a substantial increase in commensal skin bacteria that promote pathology and enhance immunity Evgeniya V Shmeleva, Mercedes Gomez de Agüero, Josef Wagner, Anton J Enright, Andrew J Macpherson, Brian J Ferguson, Geoffrey L Smith PLoS pathogens 18 (4), e1009854 |
2022 |
| Selective modulation of cell surface proteins during vaccinia infection: A resource for identifying viral immune evasion strategies Delphine M Depierreux, Arwen F Altenburg, Lior Soday, Alice Fletcher-Etherington, Robin Antrobus, Brian J Ferguson, Michael P Weekes, Geoffrey L Smith PLoS Pathogens 18 (6), e1010612 |
2022 |
| Chicken cGAS senses fowlpox virus infection and regulates macrophage effector functions Marisa Oliveira, Damaris Ribeiro Rodrigues, Vanaique Guillory, Emmanuel Kut, Efstathios S Giotis, Michael A Skinner, Rodrigo Guabiraba, Clare E Bryant, Brian J Ferguson Frontiers in Immunology 11, 3716 |
2021 |
| Vaccinia Virus Infection Inhibits Skin Dendritic Cell Migration to the Draining Lymph Node Juliana Bernardi Aggio, Veronika Krmeská, Brian J Ferguson, Pryscilla Fanini Wowk, Antonio Gigliotti Rothfuchs The Journal of Immunology 206 (4), 776-784 |
2021 |
| Exploiting Connections for Viral Replication Louise H Wong, James R Edgar, Andrea Martello, Brian J Ferguson, Emily R Eden Frontiers in Cell and Developmental Biology 9, 284 |
2021 |
| Viral interactions with non-homologous end-joining: a game of hide-and-seek Dayana B Hristova, Katharina B Lauer, Brian J Ferguson The Journal of General Virology 101 (11), 1133 |
2020 |
| ISG15-Induced IL-10 Is a Novel Anti-Inflammatory Myeloid Axis Disrupted during Active Tuberculosis Paula Fernandes dos Santos, Johan Van Weyenbergh, Murilo Delgobo, Daniel de Oliveira Patricio, Brian J. Ferguson, Rodrigo Guabiraba, Tim Dierckx, Soraya Maria Menezes, André Báfica and Daniel Santos Mansur Journal of Immunology, Volume 200(2) |
2018 |
| The Non-Homologous End Joining Protein PAXX Acts to Restrict HSV-1 Infection Ben J. Trigg, Katharina B. Lauer, Paula Fernandes dos Santos, Heather Coleman, Gabriel Balmus, Daniel S. Mansur and Brian J. Ferguson Viruses, Volume 9(11) |
2017 |
| Genome-wide analyses reveal a highly conserved Dengue virus envelope peptide which is critical for virus viability and antigenic in humans RC Fleith, FP Lobo, PF dos Santos, MM Rocha, J Bordignon, DM Strottmann, DO Patricio, WR Pavanelli, M Lo Sarzi, CN Dos Santos, BJ Ferguson, DS Mansur Scientific Reports 6, Article number: 36339 |
2016 |
| LUBAC deficiency perturbs TLR3 signaling to cause immunodeficiency and autoinflammation J Zinngrebe, E Rieser, L Taraborrelli, N Peltzer, T Hartwig, H Ren, I Kovács, C Endres, P Draber, M Darding, S von Karstedt, J Lemke, B Dome, M Bergmann , BJ Ferguson*, H Walczak* J Experimental Medicine (In Press) |
2016 |
| Increased attenuation but decreased immunogenicity by deletion of multiple vaccinia virus immunomodulators RP Sumner, H Ren, BJ Ferguson, GL Smith Vaccine Volume 34, Pages 4827–4834 |
2016 |
| The Schistosoma mansoni T2 ribonuclease omega-1 modulates inflammasome-dependent IL-1beta secretion in macrophages BJ Ferguson, S Newland, SE Gibbs, P Tourlomousis, P Fernandes dos Santos, MN Patel, SW Hall, H Walczak, G Schramm, H Haas, DW Dunne, A Cooke, and P Zaccone International Journal for Parasitology Volume 45, Issue 13 Pages 809–813 |
2015 |
| Functions of DNA damage machinery in the innate immune response to DNA virus infection BJ Trigg and BJ Ferguson Current Opinion in Virology Volume 15, Pages 56–62 |
2015 |
| International Union of Basic and Clinical Pharmacology. XCVI. Pattern Recognition Receptors in Health and Disease CE Bryant, S Orr, BJ Ferguson, MF Symmons, JP Boyle, and TP Monie Pharmacological Reviews vol. 67 no. 2 462-504 |
2015 |
| Enhancement of CD8+ T‐cell memory by removal of a vaccinia virus NF‐κB inhibitor H Ren, BJ Ferguson, C Maluquer de Motes, RP Sumner, L Harman, GL Smith Immunology 145 (1), 34-49 |
2015 |
| The role of shed placental DNA in the systemic inflammatory syndrome of pre-eclampsia JDR Hartley, BJ Ferguson, A Moffett American Journal of Obstetrics and Gynecology Volume 213, Issue 3, Pages 268–277 |
2015 |
| Intracellular sensing of viral DNA by the innate immune system DS Mansur, GL Smith, BJ Ferguson Microbes and Infection 16 (12), 1002-1012 |
2014 |
| Stimulation of Cytoplasmic DNA Sensing Pathways In Vitro and In Vivo CH Ku, BJ Ferguson J Vis Expts e51593-e51593 |
2014 |
| A mechanism for the inhibition of DNA-PK mediated DNA sensing by a virus NE Peters, BJ Ferguson, M Mazzon, AS Fahy, E Krystofinska, R Arribas-Bosacoma, LH Pearl, H Ren, GL Smith PloS Pathogens 9(10): e1003649 |
2013 |
| Vaccinia virus immune evasion: mechanisms, virulence and immunogenicity GL Smith, CTO Benfield, CM de Motes, M Mazzon, SWJ Ember, BJ Ferguson, RP Sumner Journal of General Virology, 94: 2367-2392 |
2013 |
| A mechanism for induction of a hypoxic response by vaccinia virus M Mazzon, NE Peters, C Loenarz, EM Krysztofinska, SWJ Ember, BJ Ferguson, GL Smith Proceedings of the National Academy of Sciences, 110 (30), 12444-12449 |
2013 |
| Vaccinia virus protein N2 is a nuclear IRF3 inhibitor that promotes virulence BJ Ferguson, CTO Benfield, H Ren, VH Lee, GL Frazer, P Strnadova, RP Sumner, GL Smith Journal of General Virology, 94 (9), 2070-2081 |
2013 |
| Poxvirus Targeting of E3 Ligase β-TrCP by Molecular Mimicry: A Mechanism to Inhibit NF-κB Activation and Promote Immune Evasion and Virulence DS Mansur, CM de Motes, L Unterholzner, RP Sumner, BJ Ferguson, H Ren, P Strnadova, AG Bowie, GL Smith PLoS pathogens 9 (2), e1003183 |
2013 |
| Vaccinia virus protein C4 inhibits NF-κB activation and promotes virus virulence SWJ Ember, H Ren, BJ Ferguson, GL Smith Journal of General Virology 93 (Pt 10), 2098-2108 |
2012 |
| DNA-PK is a DNA sensor for IRF-3-dependent innate immunity BJ Ferguson, DS Mansur, NE Peters, H Ren, GL Smith eLife Sciences 1 e00047 |
2012 |


